
NEWS Friday May 22-2026
2023-03-30
2023-01-24
2023-01-06
2023-01-06
2023-01-06
2023-01-06
2023-01-06
2021-05-28
2021-05-28
2021-05-28
2021-05-27
FOCUS OF THE LAB
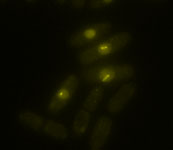
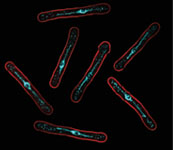


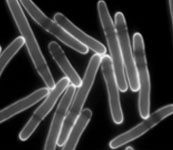
Our lab is taking a synthetic biology approach to this question, focusing on the in vivo implementation of functional model circuits to understand the basis of natural cellular networks. We use the fission yeast cell cycle as a model eukaryotic system to understand: 1) the core molecular and dynamic inputs that control the sequence of cell cycle events and that limit the variability in cell cycle execution; 2) the establishment of optimal cell proliferation through evolutionary mechanisms; and 3) why evolution has chosen to adopt complexity in certain regulatory networks when it may appear to be dispensable. In the same way that model organisms have been pivotal for revealing the operation of conserved mechanisms, synthetic biology will allow us to extract general principles that govern the organization and evolution of essential cellular functions.

